
DNA Encoded Library Software.
Industry standard software for DEL hit picking, resynthesis prioritisation, and virtual screening from DEL data.

Industry standard software for DEL hit picking, resynthesis prioritisation, and virtual screening from DEL data.






Cambridge Molecular officially announces a major update to its Palladium DEL Follow-Up Suite. The Palladium DEL Follow-Up Suite is a fully-featured downloadable offline software application for handling DEL datasets… Read More

Palladium is a downloadable software for DNA encoded library data analysis, virtual screening, and resynthesis prioritisation. Palladium is compatible with all DEL builds and manufacturers, is available on all major operating systems, and comes with full data analysis support.

Palladium DEL Follow-Up Suite DNA Encoded Library Software includes:
A sophisticated shape and pharmacophore consensus discovery algorithm that looks for scaffold-hopping networks within a DEL dataset. Enables quality triage of DEL datasets, resynthesis prioritisation, and 3D virtual screening setup.
A fast and effective 2D virtual screening system that accepts entire raw DEL datasets as input. DELNexus is capable of both broad-net and focussed virtual screens, and automatically leverages the full computational resources available.
A lightning-fast 3D virtual screening system that accepts PocketMapper models or ligand crystal structures as input. ShapeRetriever can screen through millions of conformers with ease, and includes a catalogue generation function.
Our fast and robust implementation of 2D and 3D DEL data plots, which includes automatic discovery and selection of enriched disynthon features.
A robust selectivity analysis tool for comparing multiple selection conditions with each other. Arbitrary regions of the plot can be lassoed and exported.


PocketMapper is a sophisticated shape and pharmacophore analysis algorithm that identifies scaffold-hopping networks within DEL datasets. Numerous useful pieces of information can be generated from a scaffold-hopping network, including:



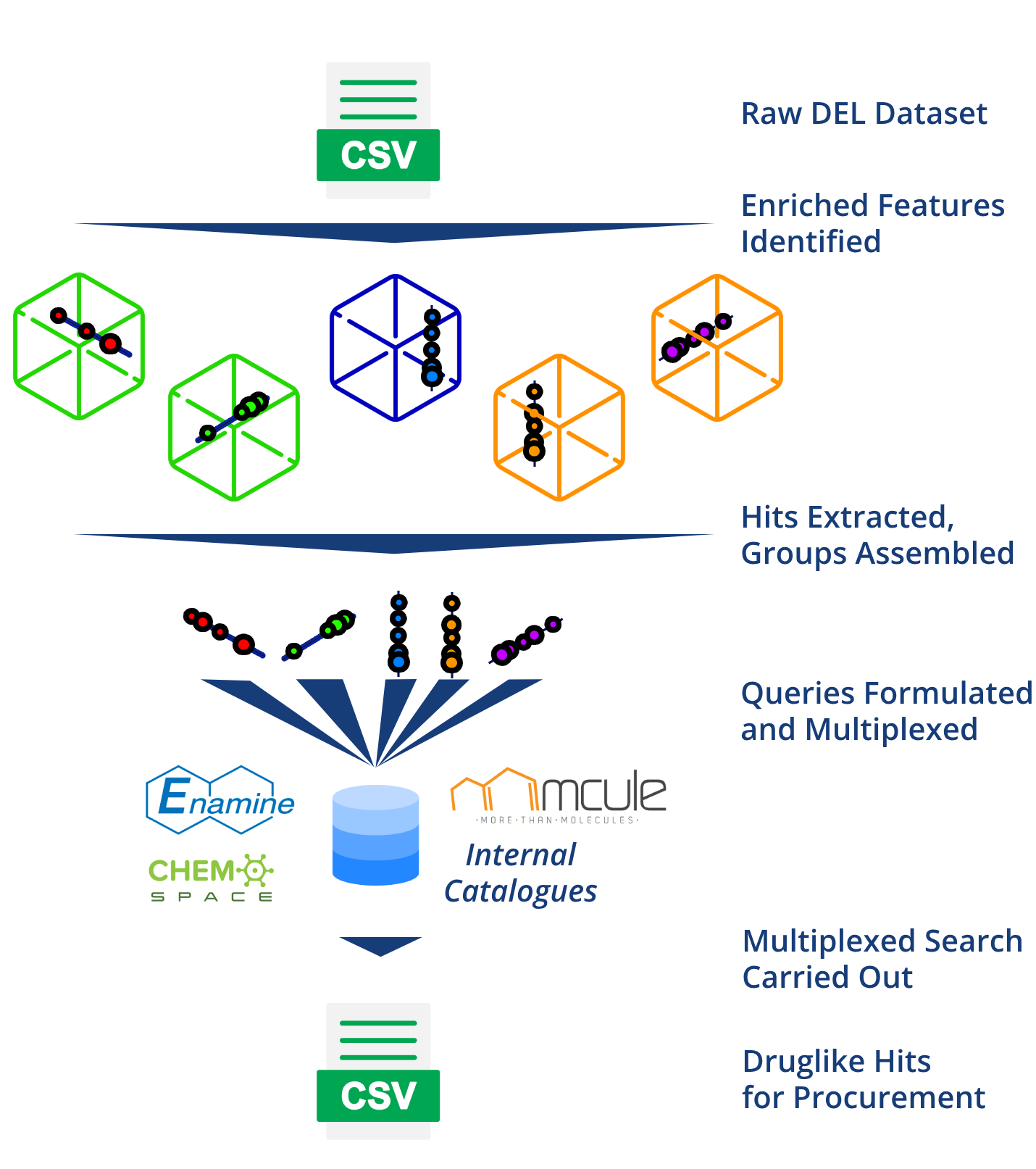
DELNexus is a rapid and precise virtual screening system, that takes an entire DEL dataset as input. Casting a broad net into chemical space, DELNexus identifies enriched groups of DEL hits from all libraries, then uses our PathFinder algorithm to identify an exhaustive list of all important substructures, and the most efficient search routine for finding catalogue compounds that contain them. The tool is the fastest and easiest way to get druglike chemical matter into your lab after a DEL selection.






ShapeRetriever uses the volume, hydrophobic volume, acceptor volume, donor volume, and occluded volume of a 3D query to search for matching compounds in a catalogue of interest. The tool is extremely fast and highly effective. Used in conjunction with PocketMapper, it can find active hits even when no crystal structure yet exists for the target.

Book a demo below.

Our team will reply as soon as they can.
